XAGE-1, a Cancer Testis Antigen: Potential Use as a Molecular Target for Ewing’s Sarcoma
XAGE-1, a Cancer Testis Antigen: Potential Use as a Molecular Target for Ewing’s Sarcoma
By Kristi A. Egland, PhD, and Tapan K. Bera, PhD
The identification of cancer antigens provides new opportunities for the development of therapeutic strategies against cancer. During the last several years, researchers have developed two major approaches to identify human tumor antigens that are recognized by cytotoxic T lymphocytes (CTL). The first approach is based on direct biochemical purification of tumor antigen peptides from the peptide-MHC complexes, which are recognized by the CTL.1 The second is a traditional genetic approach to identify tumor antigens; it relies on the generation of a genomic or cDNA library from the tumor cell line. The DNA library is transfected into cells expressing the appropriate MHC molecule. Cells that present peptides derived from proteins encoded by the cDNA library are isolated on the basis of the ability to stimulate cytokine release from CTL.2 Melanoma antigen (MAGE-1) was the first tumor antigen identified from a human melanoma using the traditional genetic approach.2
Subsequently, several cancer antigens have been identified using other approaches, and some of these antigens have been recognized as candidates for cancer vaccines.3 MAGE-1 is a member of the family of cancer testis (CT) antigens, which is the best characterized group of antigens so far. CT antigens are a distinct class of antigens that are expressed in many types of cancers, yet they have a restricted expression pattern in normal tissues.4-6
These genes primarily are expressed in the primitive germ cells, spermotogonia, and normal testis.7 Some extensively studied CT antigens are MAGE, GAGE, BAGE, and GnT-V and their family members. Although the previously mentioned CT antigens were identified from cDNA libraries derived from a single melanoma tumor cell line, malignant transformation often is associated with activation or depression of silent CT antigens in a wide range of human tumors.3
EST Database Mining and Identification of Novel Cancer Antigens
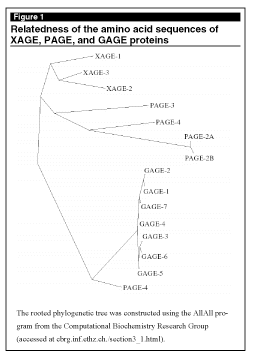
Our laboratory is using a functional genomics approach to discover new genes that are associated with cancer.8 The cancer genome anatomy project of the National Cancer Institute uses laser-captured microdisection techniques to generate EST libraries. ESTs are partial sequences of cDNA clones randomly selected from various cDNA libraries. Because each of these clones is generated from a single transcript, the number of ESTs for a particular gene from the same library provides valuable information on the expression patterns of genes in different tissues. These EST sequences can be clustered and sorted to identify genes that are preferentially or exclusively expressed in malignant tissues. We have reported a computer screening strategy to identify genes that are preferentially expressed in prostate tumors. By using this screen, we have identified several novel genes, including PAGE-4, which is homologous to the GAGE family of cancer testis antigens.8,9 To discover additional new tumor antigens, we used a "homology walking" program and identified three novel PAGE-GAGE-related genes, including XAGE-1. (See Figure 1.)
 |
XAGE Expression in Different Tissues
The computer-based screening strategy developed by our group lead to the identification of the novel gene XAGE-1.9 The XAGE-1 cluster contains ESTs from testis, alveolar rhabdomyosarcoma, Ewing’s sarcoma, and germ cell tumor cDNA libraries. (See Figure 2.) The XAGE-1 expression pattern in normal tissues was experimentally determined.10 A RNA dot blot analysis revealed that XAGE-1 is expressed in normal testis. A reverse transcriptase-polymerase chain reaction (RT-PCR) analysis, using a human rapid-scan panel containing cDNAs from 24 different tissues of the body, confirmed the high expression of XAGE-1 in normal testis, but also showed that XAGE-1 was expressed at low levels in lung and peripheral blood lymphocytes (PBL).
 |
However, Northern blot analysis, using a fragment of XAGE-1 as a probe, revealed a single transcript of approximately 700 bp in testis; no signal was detected in lung or PBL. (See Figure 3.) Because Northern blot analysis is less sensitive than RT-PCR, the above results all consistently showed that XAGE-1 is abundantly expressed in the normal testis only, and weakly expressed in lung and PBL. Based on sequence similarity and expression pattern, XAGE-1 is a member of the family of CT antigens. The XAGE-1 gene is located on the X chromosome,10 which is consistent with the location of other CT antigen genes.12
 |
XAGE Expression in Ewing’s Sarcoma and Other Cancers
The Ewing’s family of tumors include: Ewing’s tumor of bone; extraosseus Ewing’s (tumor growing outside of the bone); primitive neuroectodermal tumor (PNET), also known as peripheral neuroepithelioma; and Askin’s tumor (PNET of the chest wall). These tumors are rare diseases in which cancer cells are found in the bone and soft tissues.13 This family of tumors has specific chromosomal translocations that result in the fusion of the EWS and FLI1 genes. The FLI1 gene encodes a member of the ETS family of transcription factors,14 while the widely expressed EWS gene encodes a protein of unknown function. The site disrupted by the chromosomal translocation found in the Ewing’s sarcoma group of tumors is located in the EWS gene.15 Chromosomal analysis of Ewing’s sarcoma has demonstrated that about in 90-95% of cases a t(11;22)(q24;q12) translocation generates an EWS-FLI1 fusion protein, and the precise exon composition of the fusion transcript has been found to be a prognostic marker in this group of tumors.14 Although the EWS-FLI1 chimeric protein may have therapeutic potential as a molecular target, other tumor-specific targets for Ewing’s sarcoma need to be identified.
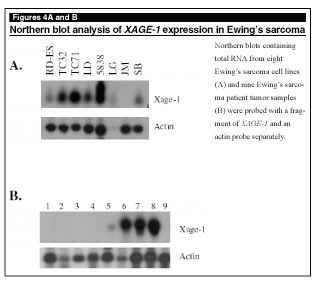
Our EST database analysis predicts that XAGE-1 is expressed in alveolar rhabdomyosarcoma and Ewing’s sarcom.9 Consistent with this prediction, XAGE-1 was expressed in one of one patient samples with alveolar rhabdomyosarcoma and one of three patient samples with embryonal rhabdomyosarcoma. In addition, the XAGE-1 transcript was present in two of five osteosarcoma cell lines.10 We also investigated the expression of XAGE-1 in Ewing’s sarcoma. Northern blot analysis revealed an XAGE-1 transcript in seven of eight Ewing’s sarcoma cell lines. (See Figure 4A.) Interestingly, the one cell line that does not express XAGE-1 also lacks the EWS-FLI-1 chromosomal translocation. XAGE-1 also was expressed in four of nine human patient Ewing’s sarcoma samples. (See Figure 4B.) We found a correlation between the presence of the EWS-FLI-1 or EWS-Erg chromosome translocation and the presence of the XAGE-1 transcript. However, XAGE-1 was not expressed in every patient sample that contains the chromosome translocation. The significance of this correlation, and the possibility that XAGE-1 expression could be under the control of the EWS-FLI-1 fusion protein, still remains to be determined.
 |
Implications
The ideal cellular target for immunotherapeutic treatment of cancer, such as cancer vaccines or immunotoxins, is one that predominantly has restricted expression in diseased tissues, with some expression in dispensable organs. By using such targets, engineered immunotherapies can kill the tumor cells while sparing the normal tissues. As of yet, good molecular targets for the Ewing’s family of tumors still need to be identified. Because XAGE-1 is expressed in testis and in a large number of Ewing’s sarcomas, as well as other cancers, it has good potential for use as a target for various kinds of immunotherapies. (Dr. Egland is Postdoctoral Fellow and Dr. Bera is Staff Scientist in the Laboratory of Molecular Biology, National Cancer Institute, National Institutes of Health, Bethesda, MD.)
Acknowledgements
We would like to thank the members of the Gene Discovery Group for their contributions to this project and Ira Pastan for his support and valuable suggestions.
References
1. Mandelboim O, Berke G, Fridkin M, et al. CTL induction by a tumor-associated antigen octapeptide derived from a murine lung-carcinoma. Nature 1994;369:67-71.
2. Van der Bruggen P, Traversari C, Chomez P, et al. A gene encoding an antigen recognized by cytolytic T-lymphocytes on a human-melanoma. Science 1991;254:1643-1647.
3. Wang RF, Rosenberg SA. Human tumor antigens recognized by T lymphocytes: Implications for cancer therapy. J Leuk Biol 1996;60:296-309.
4. De Smet C, Lurquin C, Plaen ED, et al. Genes coding for melanoma antigens recognized by cytolytic lymphocytes. Eye 1997;11:243-248.
5. Chen YT, Old LJ. Cancer-testis antigens: Targets for cancer immunotherapy. Cancer J Sci Am 1999;5:16-17.
6. Gillespie AM, Rodgers S, Wilson AP, et al. MAGE, BAGE and GAGE: Tumor antigen expression in benign and malignant ovarian tissue. Br J Cancer 1998;78: 816-821.
7. Takahashi K, Shichijo S, Noguchi M, et al. Identification of MAGE-1 and MAGE-4 proteins in spermatogonia and primary spermatocytes of testis. Cancer Res 1995;55:3478-3482.
8. Vasmatzis G, Essand M, Brinkmann U, et al. Discovery of three genes specifically expressed in human prostate by expressed sequence tag database analysis. Proc Natl Acad Sci U S A 1998;95:300-304.
9. Brinkmann U, Vasmatzis G, Lee B, et al. Novel genes in the PAGE and GAGE family of tumor antigens found by homology walking in the dbEST. Cancer Res 1999;59:1445-1448.
10. Liu XF, Helman LJ, Yeung C, et al. XAGE-1, A new gene that is frequently expressed in Ewing’s sarcoma. Cancer Res 2000;60:4752-4755.
11. De Plaen E, Arden K, Traversari C, et al. Structure, chromosomal localization, and expression of 12 genes of the MAGE family. Immunogenetics 1994;40: 360-369.
12. Rogner UC, Wilke K, Steck E, et al. The melanoma antigen gene (MAGE) family is clustered in the chromosomal band Xq28. Genomics 1995;29:725-731.
13. de Alava E, Gerald WL. Molecular biology of the Ewing’s sarcoma/primitive neuroectodermal tumor family. J Clin Oncol 2000;18:204-213.
14. Delattre O, Zucman J, Plougastel B, et al. Gene fusion with an ETS DNA-binding domain caused by chromosome-translocation in human tumors. Nature 1992; 359:162-165.
15. Ohno T, Ouchida M, Lee L, et al. The EWS gene, involved in Ewing family of tumors, malignant-melanoma of soft parts and desmoplastic small round-cell tumors, codes for an RNA-binding protein with novel regulatory domains. Oncogene 1994;9:3087-3097.
Subscribe Now for Access
You have reached your article limit for the month. We hope you found our articles both enjoyable and insightful. For information on new subscriptions, product trials, alternative billing arrangements or group and site discounts please call 800-688-2421. We look forward to having you as a long-term member of the Relias Media community.
