Something in Common Between Candida albicans and Staphylococcus aureus: Coagulase Production
Something in Common Between Candida albicans and Staphylococcus aureus: Coagulase Production
Abstract & Commentary
Synopsis: Candida albicans and some other Candida species elaborate coagulase and might be confused with Staphylococcus aureus.
Source: Rodrigues AG, et al. Expression of plasma coagulase among pathogenic Candida species. J Clin Microbiol. 2003; 41:5792-5793.
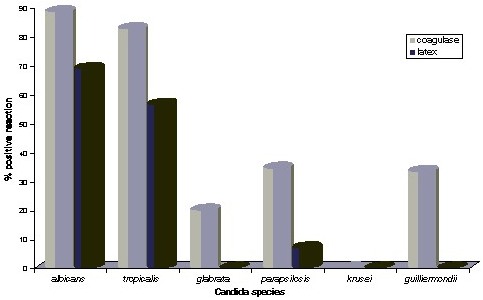
A variety of different clinical Candida isolates totaling 161 from blood, respiratory secretions, genital secretions, stool, and urine were tested for the coagulase reaction using the classical plasma test, as well as a commercial latex test (Pastorex Staph-Plus kit [Bio-Rad]). The identity of the different species (C albicans [70 isolates]; C tropicalis [23 isolates]; C glabrata [25 isolates]; C parapsilosis [29 isolates]; C krusei [11 isolates]; and C guilliermondii [3 isolates]) was confirmed by the germ tube test and the API 32C identification kit (BioMérieux). Coagulase production was indicated by clot formation after 4 h in rabbit plasma or, if negative, 20 h later. The latex particles in the Staph-Plus kit are sensitized with human fibrinogen and monoclonal antibodies and react with the clumping factor, protein A, and capsular polysaccharides of Staphylococcus aureus. Most of the C albicans (88.5%) and C tropicalis (82.6%) strains were able to induce clot formation from EDTA-rabbit plasma at 24 hours and react with the latex test (see Figure). The other species reacted to a much lesser extent or not at all. Rodrigues and colleagues felt that this was more than a curiosity, especially in the setting of "a large university hospital, handling a considerable number of samples daily and relying mostly on automated devices for identity confirmation" because yeasts might be taken for S aureus when the latex coagulase test is performed to identify white colonies appearing on agar plates after overnight incubation unless microscopic examination is done.
|
Coagulase Reactions Among Candida Species |
|
|
Comment by J. Peter Donnelly, PhD
I confess I was surprised to learn that C albicans and S aureus had something in common after all. But, then again, I was brought up on steam-age microbiology in which one would not dream of proceeding from plate to test without first carefully noting the size, shape, color, and texture of any colonies formed on petri dishes; recording the time, temperature and atmosphere of incubation; and establishing the Gram reaction and shape, and the catalase and oxidase reactions. Of course, shortcuts could, and would, be taken, but there was a safeguard that final reports would only be issued when all the results were known. This, of course, took timetoo much time in many cases. Rightly so, we have come to expect rapid results, economic use of resources, and short turn-around times. However, as shown in this article, micro-organisms continue to baffle and fool us, especially when we have taken our eye off the ball. Nowadays, it is perfectly possible to pick off a colony and surrender it entirely to an electronic black box and have the result within a few hours. It is only a short step in thrift to eliminate skilled personnel altogether from this process. Many professionals’ worst nightmare is that we might end up using systems beyond our control, employing techniques we don’t fully understand, and releasing results we cannot rely on. How many of us have deliberately put the wrong organism in an identification kit to educate students and tease colleagues? There was also a serious sidegetting the message across that elementary microbiology is and will remain our best safeguard against making mistakes.
Dr. Donnelly, Clinical Microbiologist, University Hospital Nijmegen, The Netherlands, is Associate Editor of Infectious Disease Alert.
Candida albicans and some other Candida species elaborate coagulase and might be confused with Staphylococcus aureus.Subscribe Now for Access
You have reached your article limit for the month. We hope you found our articles both enjoyable and insightful. For information on new subscriptions, product trials, alternative billing arrangements or group and site discounts please call 800-688-2421. We look forward to having you as a long-term member of the Relias Media community.